
Plotting outputs
Source:R/ArraySpeciesBySize-class.R, R/ArrayTimeBySpecies-class.R, R/ArrayTimeBySpeciesBySize-class.R
plot.RdMany mizer functions return values that depend on species and either size or
time. plot() creates a ggplot2 figure with one line for each species
showing the values against size or against time (depending on the type of
output). ggplotly() creates an interactive version of the same figure.
Usage
# S3 method for class 'ArraySpeciesBySize'
plot(
x,
species = NULL,
all.sizes = FALSE,
highlight = NULL,
return_data = FALSE,
log_x = TRUE,
log_y = FALSE,
log = NULL,
wlim = c(NA, NA),
ylim = c(NA, NA),
total = FALSE,
background = TRUE,
y_ticks = 6,
...
)
# S3 method for class 'ArraySpeciesBySize'
ggplotly(x, ...)
# S3 method for class 'ArrayTimeBySpecies'
plot(
x,
species = NULL,
start_time = NULL,
end_time = NULL,
y_ticks = 6,
ylim = c(NA, NA),
total = FALSE,
background = TRUE,
highlight = NULL,
log_x = FALSE,
log_y = TRUE,
log = NULL,
return_data = FALSE,
...
)
# S3 method for class 'ArrayTimeBySpecies'
ggplotly(x, ...)
# S3 method for class 'ArrayTimeBySpeciesBySize'
plot(
x,
species = NULL,
time = NULL,
all.sizes = FALSE,
highlight = NULL,
return_data = FALSE,
log_x = TRUE,
log_y = FALSE,
log = NULL,
wlim = c(NA, NA),
ylim = c(NA, NA),
total = FALSE,
background = TRUE,
y_ticks = 6,
...
)
# S3 method for class 'ArrayTimeBySpeciesBySize'
ggplotly(x, ...)Arguments
- x
An
ArraySpeciesBySizeorArrayTimeBySpeciesobject.- species
Character vector of species to include.
NULL(default) means all species.- all.sizes
If
FALSE(default), values outside a species' size range (w_mintow_max) are removed. Only applies toArraySpeciesBySize.- highlight
Name or vector of names of the species to be highlighted.
- return_data
If
TRUE, return the data frame instead of the plot.- log_x
If
TRUE, use a log10 x-axis. Default isTRUEfor size spectra andFALSEfor time series.- log_y
If
TRUE, use a log10 y-axis. Default isFALSEforArraySpeciesBySizeandTRUEforArrayTimeBySpecies.- log
Character string specifying which axes should use log10 scales, in the same form as the base
plot()argument. For example,"x","y","xy"or"". If supplied, this overrideslog_xandlog_y.- wlim
A numeric vector of length two providing lower and upper limits for the weight (x) axis. Use
NAto refer to the existing minimum or maximum. Only applies toArraySpeciesBySize.- ylim
A numeric vector of length two providing lower and upper limits for the value (y) axis. Use
NAto refer to the existing minimum or maximum.- total
A boolean value that determines whether the total over all selected species is plotted as well. Default is
FALSE.- background
A boolean value that determines whether background species are included. Ignored if the model does not contain background species. Default is
TRUE.- y_ticks
The approximate number of ticks desired on the y axis.
- ...
Further arguments (currently unused).
- start_time
The first time to be plotted. Default (
NULL) is the beginning of the time series. Only applies toArrayTimeBySpecies.- end_time
The last time to be plotted. Default (
NULL) is the end of the time series. Only applies toArrayTimeBySpecies.- time
The time to display. Default (
NULL) is the final time step. Only applies toArrayTimeBySpeciesBySize.
Value
plot() returns a ggplot2 object, unless return_data = TRUE, in
which case a data frame is returned.
ggplotly() returns a plotly object.
Details
This works because the mizer functions that give values that depend on
species and size return an ArraySpeciesBySize object and those that
give values that depend on species and time return an ArrayTimeBySpecies
object. These objects have attributes that store the name of the value,
its units, and a reference to the MizerParams object that the value was
computed from. This allows the plots to be automatically labelled and
coloured appropriately.
See also
Other plotting functions:
addPlot(),
animate.ArrayTimeBySpeciesBySize(),
plot2(),
plotBiomass(),
plotCDF(),
plotCDF2(),
plotDiet(),
plotFMort(),
plotFeedingLevel(),
plotGrowthCurves(),
plotMizerParams,
plotMizerSim,
plotPredMort(),
plotRelative(),
plotSpectra(),
plotSpectra2(),
plotSpectraRelative(),
plotYield(),
plotYieldGear(),
plotting_functions
Examples
# \donttest{
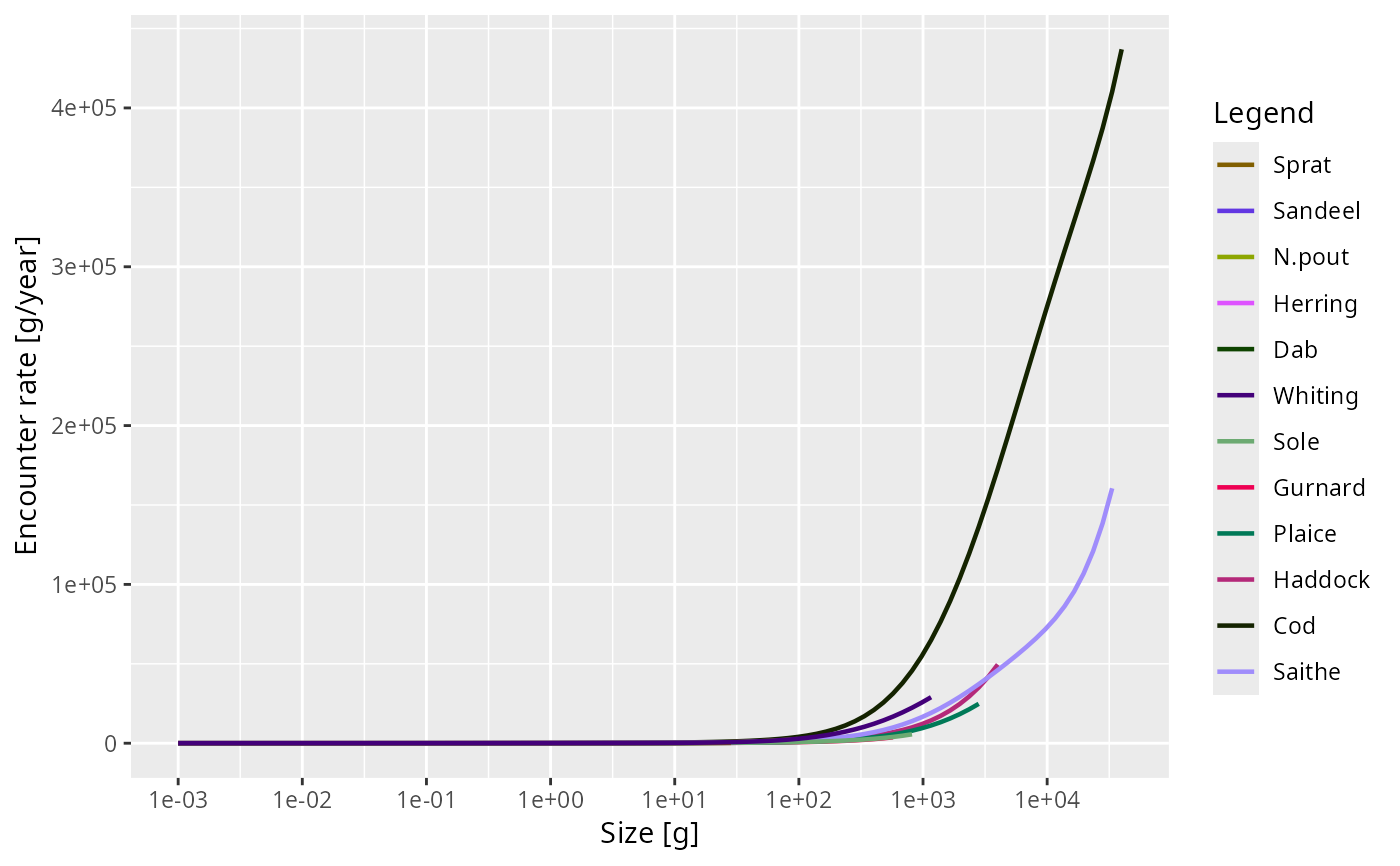
plot(getEncounter(NS_params))
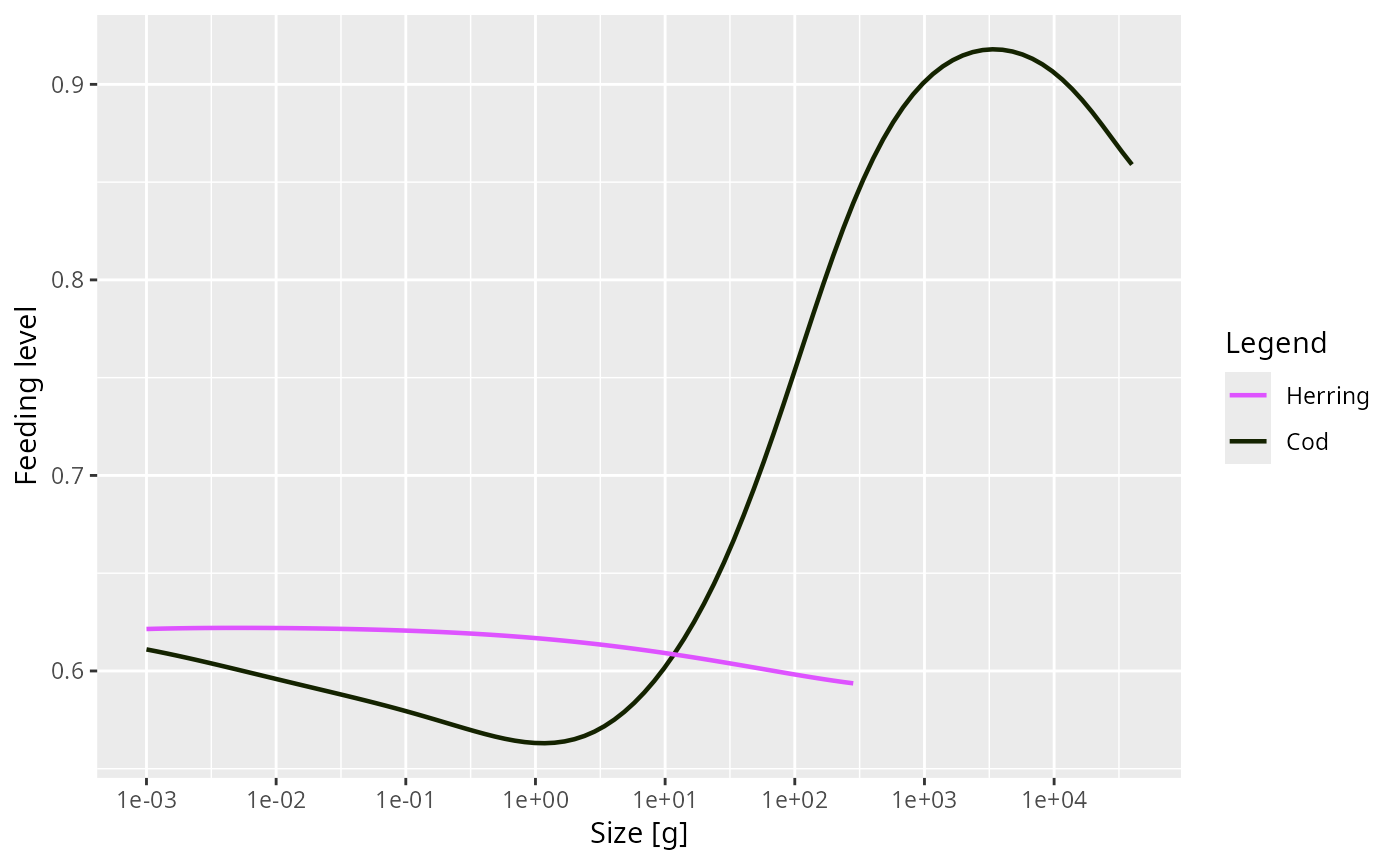
 plot(getFeedingLevel(NS_params), species = c("Cod", "Herring"))
plot(getFeedingLevel(NS_params), species = c("Cod", "Herring"))
 # }
# \donttest{
ggplotly(getEncounter(NS_params))
# }
# \donttest{
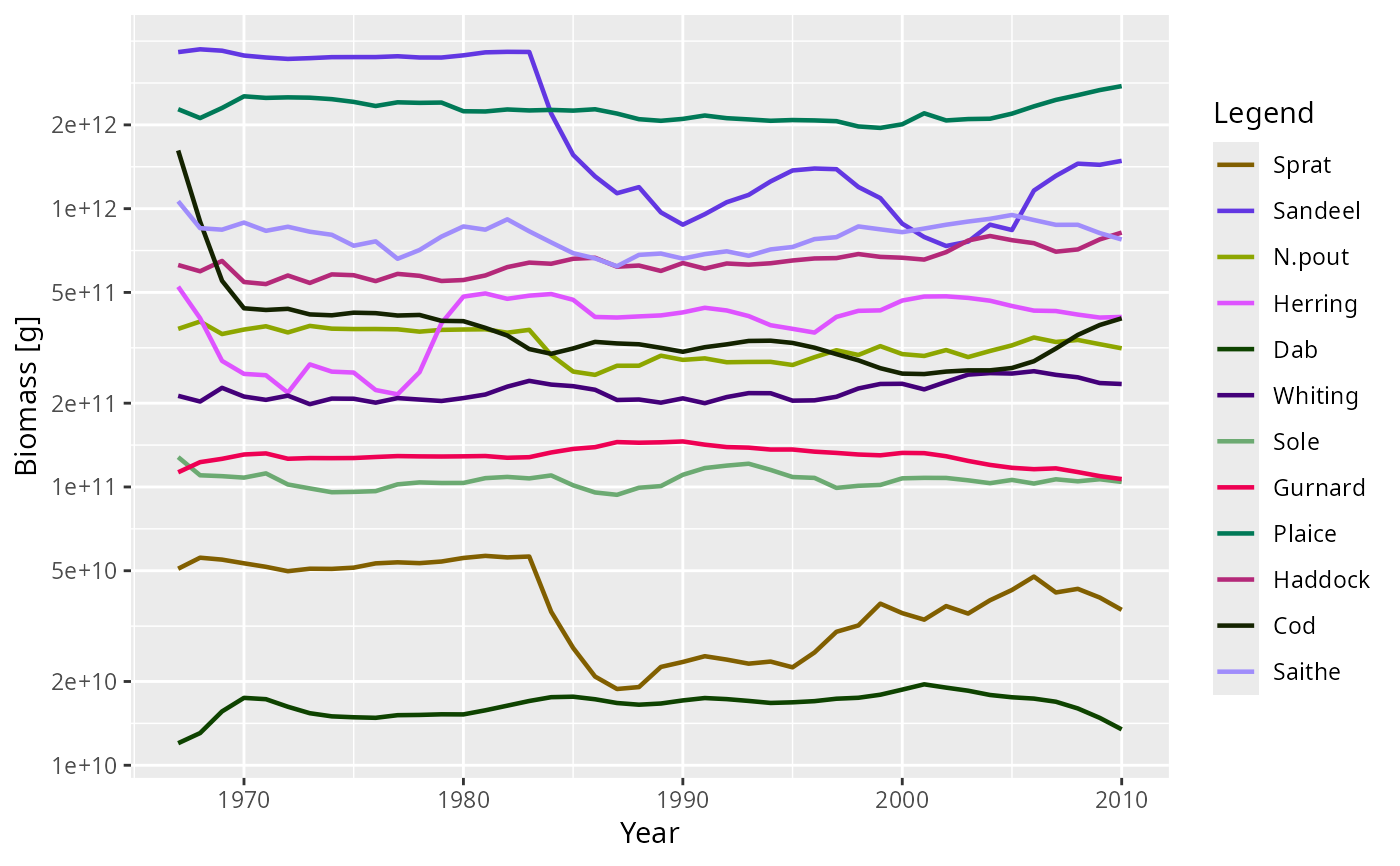
plot(getBiomass(NS_sim))
# }
# \donttest{
ggplotly(getEncounter(NS_params))
# }
# \donttest{
plot(getBiomass(NS_sim))
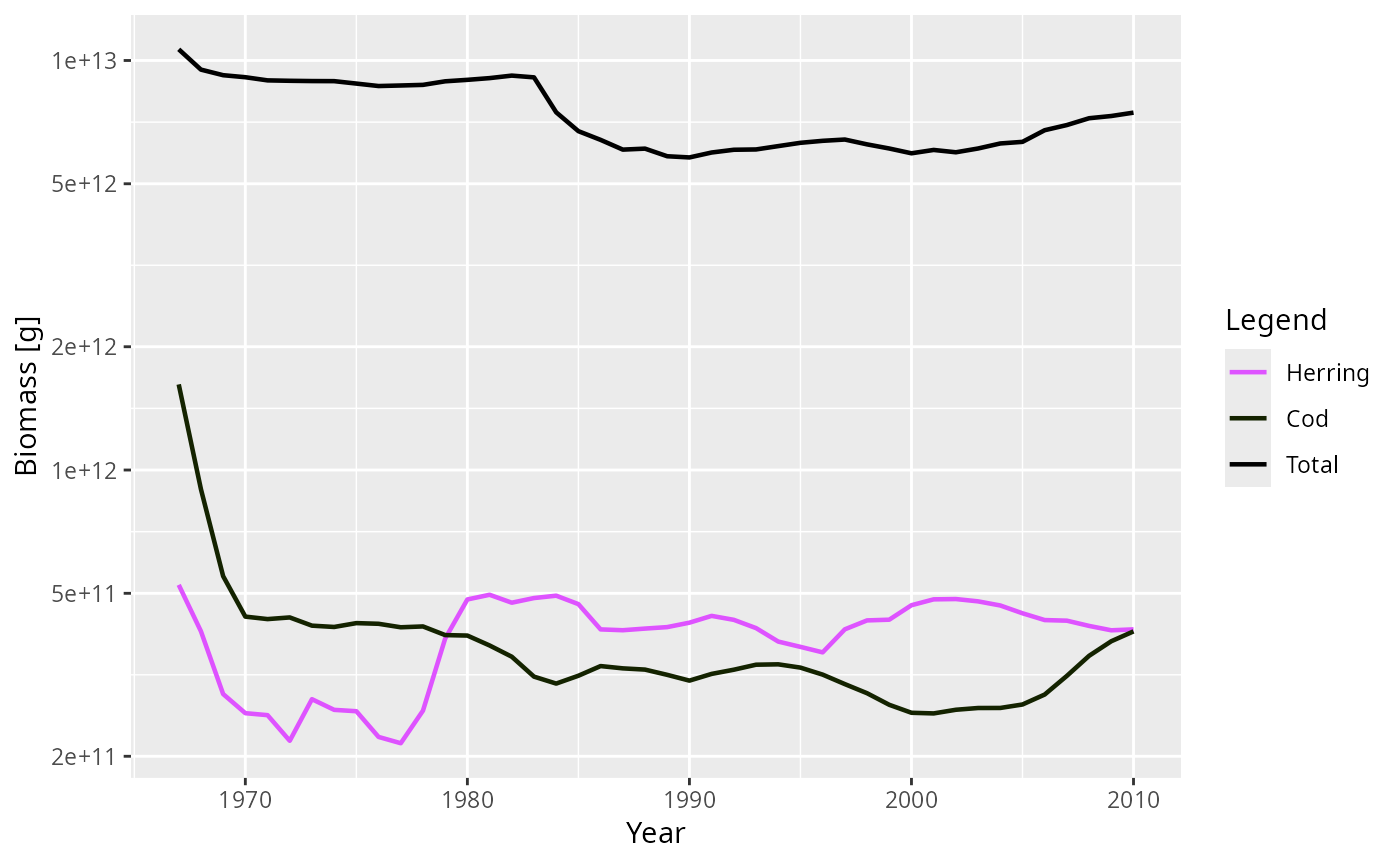
 plot(getBiomass(NS_sim), species = c("Cod", "Herring"), total = TRUE)
plot(getBiomass(NS_sim), species = c("Cod", "Herring"), total = TRUE)
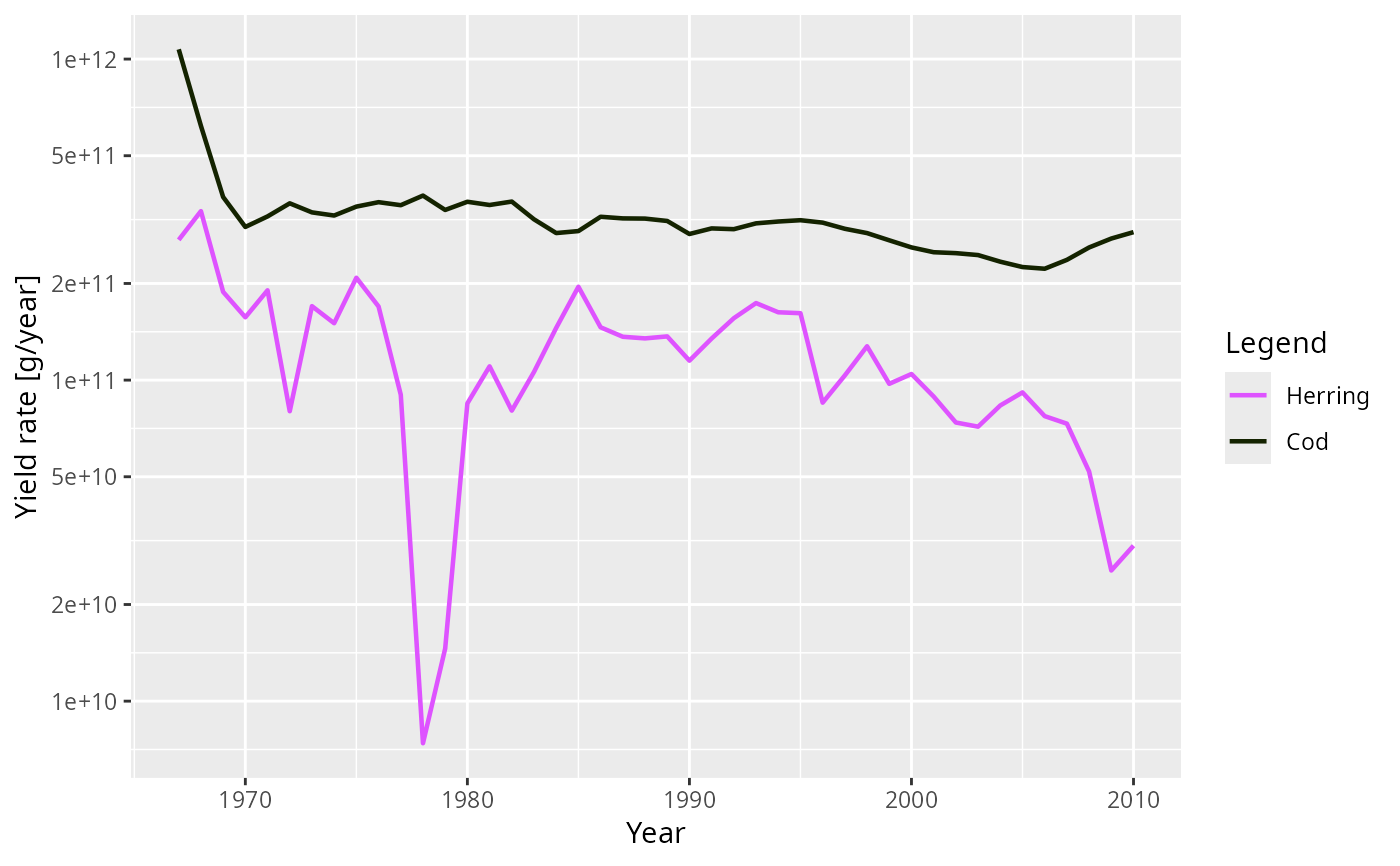
 plot(getYield(NS_sim), species = c("Cod", "Herring"))
plot(getYield(NS_sim), species = c("Cod", "Herring"))
 # }
# \donttest{
ggplotly(getBiomass(NS_sim))
# }
# \donttest{
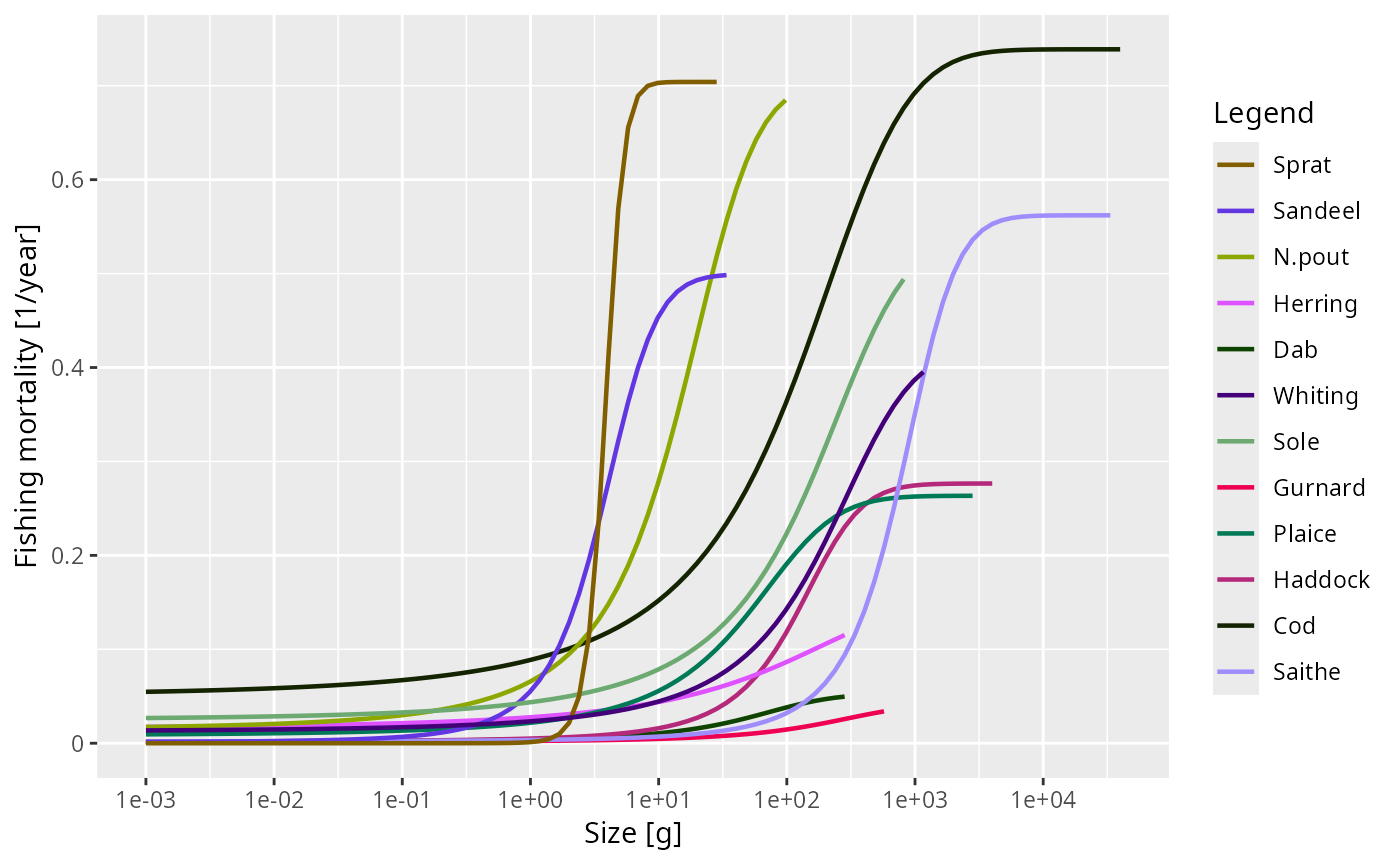
plot(getFMort(NS_sim))
# }
# \donttest{
ggplotly(getBiomass(NS_sim))
# }
# \donttest{
plot(getFMort(NS_sim))
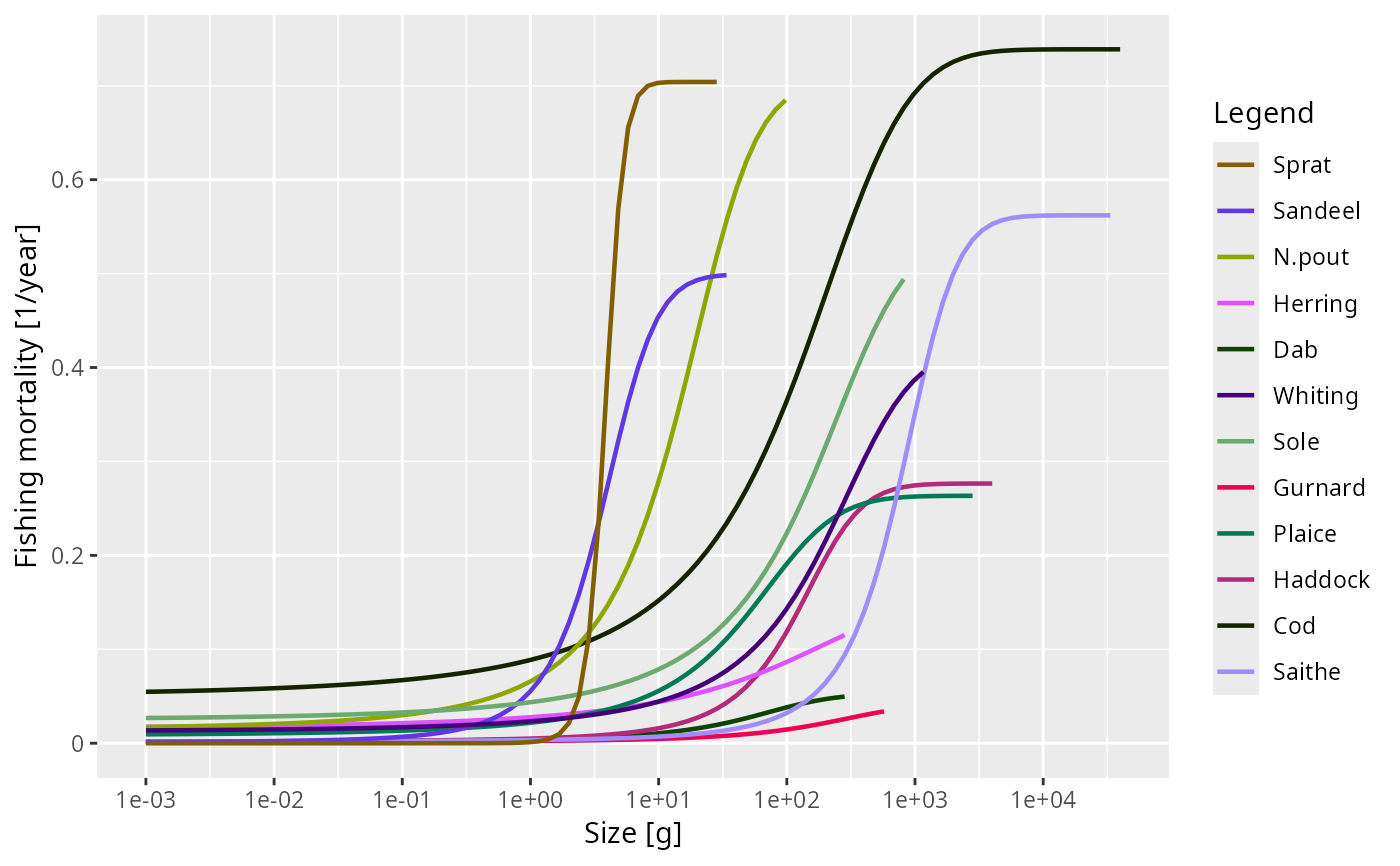
 plot(getFMort(NS_sim), time = 2010)
plot(getFMort(NS_sim), time = 2010)
 # }
# \donttest{
ggplotly(getFMort(NS_sim))
# }
# }
# \donttest{
ggplotly(getFMort(NS_sim))
# }