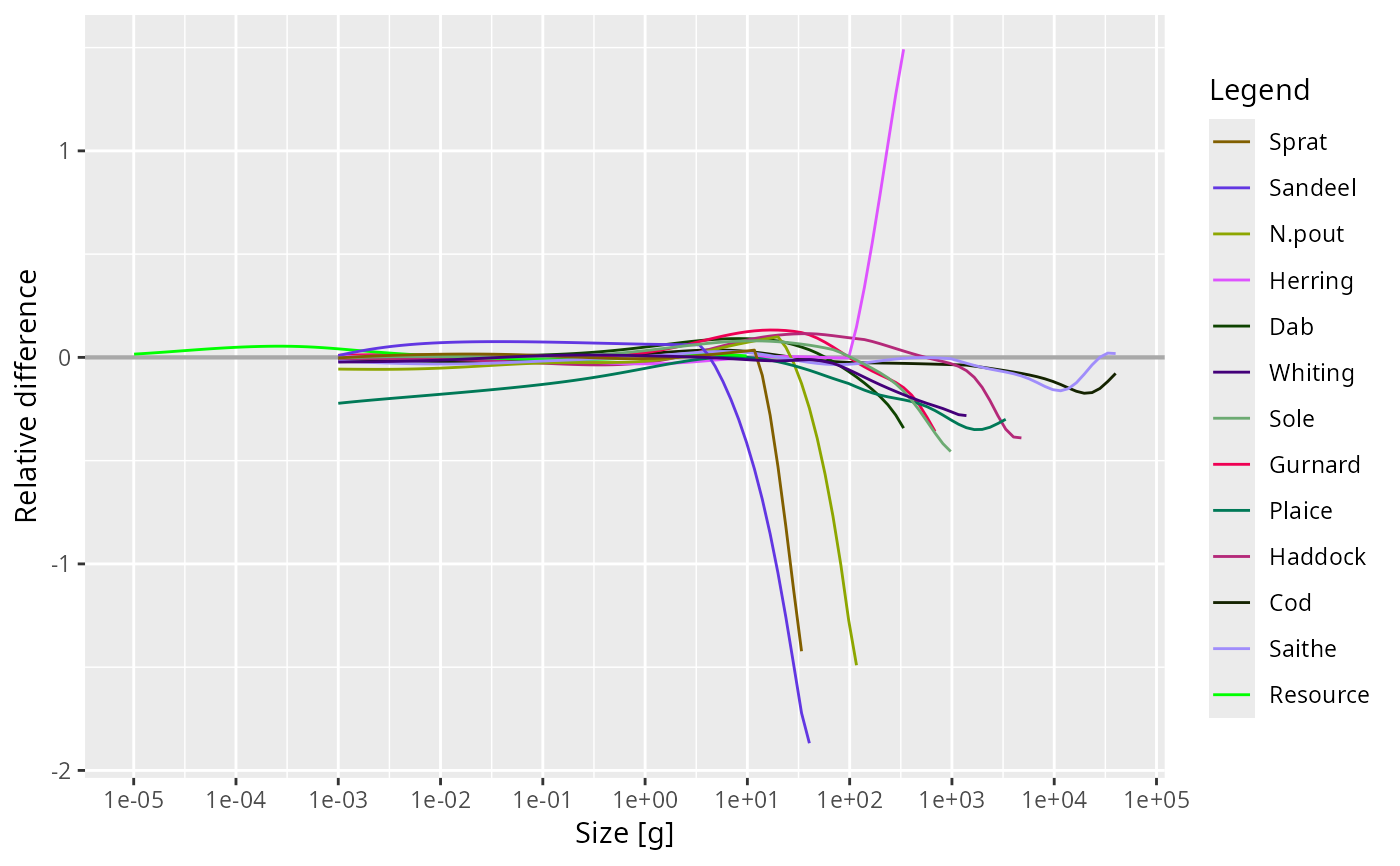
plotSpectraRelative() plots the difference between the spectra relative to
their average. If we denote the number density from the first object as
\(N_1(w)\) and that from the second object as \(N_2(w)\), then this plot
shows
$$2 (N_2(w) - N_1(w)) / (N_2(w) + N_1(w)).$$
Arguments
- object1
First
MizerParamsorMizerSimobject.- object2
Second
MizerParamsorMizerSimobject.- log_x
If
TRUE(default), use a log10 x-axis.- ylim
A numeric vector of length two providing lower and upper limits for the relative difference (y) axis. Use
NAto refer to the existing minimum or maximum.- ...
Arguments passed to
plotSpectra()for preparing the spectra data, for examplespecies,time_range,wlim,resource,backgroundortotal.
Details
The individual spectra are calculated by plotSpectra(), to which all
additional arguments are passed. For example, you can determine a time range
over which to average simulation results via time_range. See
plotSpectra() for more options.
Note that it does not matter whether the relative difference is calculated for number density, biomass density, or biomass density in log weight, because the factors of \(w\) by which the densities differ cancel out in the relative difference.
See also
Other plotting functions:
addPlot(),
animate.ArrayTimeBySpeciesBySize(),
plot,
plot2(),
plotBiomass(),
plotCDF(),
plotCDF2(),
plotDiet(),
plotFMort(),
plotFeedingLevel(),
plotGrowthCurves(),
plotMizerParams,
plotMizerSim,
plotPredMort(),
plotRelative(),
plotSpectra(),
plotSpectra2(),
plotYield(),
plotYieldGear(),
plotting_functions