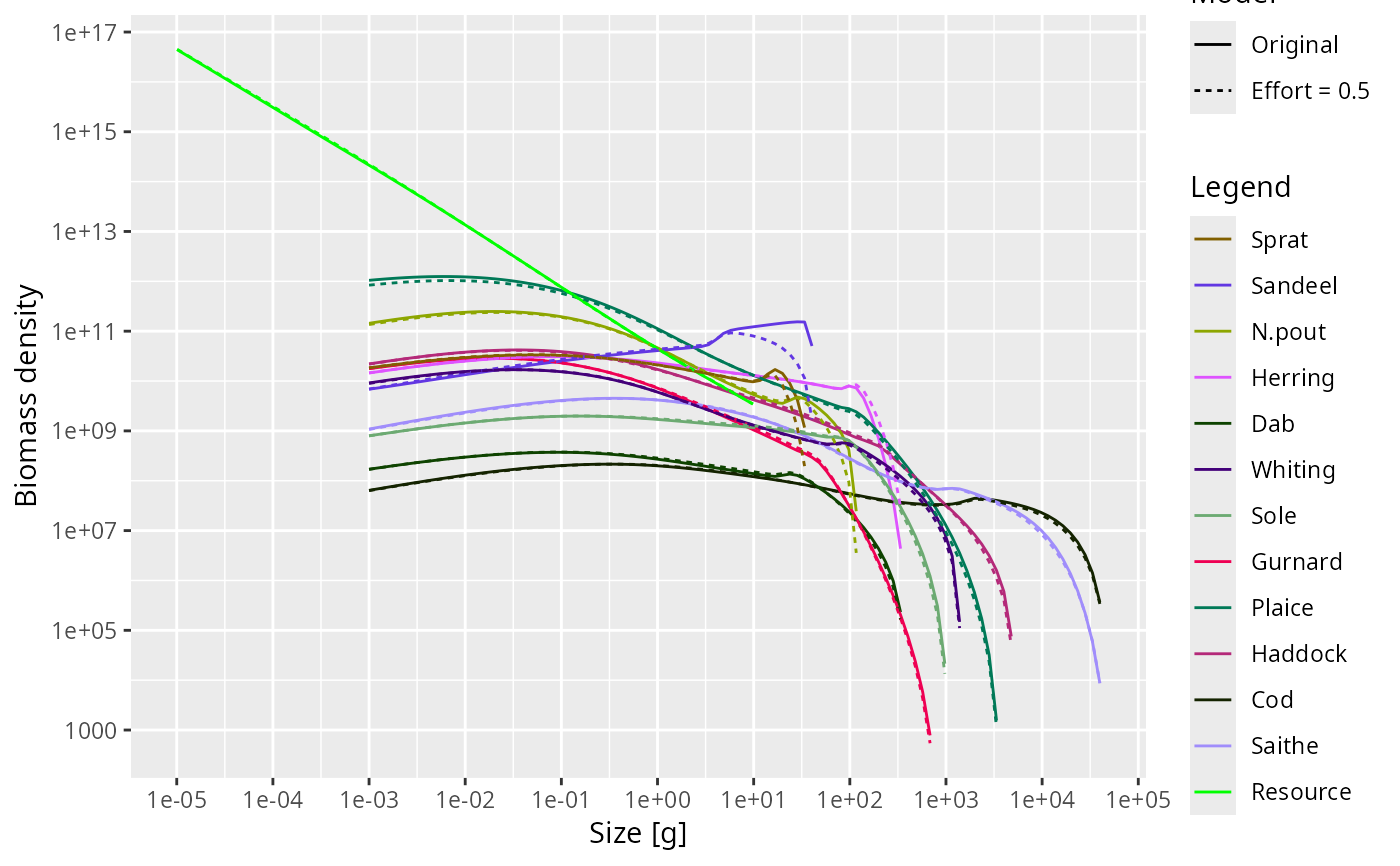
plotSpectra2() compares the abundance spectra from two MizerParams or
MizerSim objects in a single plot. Colours identify species or groups and
linetype identifies the object.
Usage
plotSpectra2(
object1,
object2,
name1 = "First",
name2 = "Second",
power = 1,
log_x = TRUE,
log_y = TRUE,
log = NULL,
...
)
plotlySpectra2(
object1,
object2,
name1 = "First",
name2 = "Second",
power = 1,
log_x = TRUE,
log_y = TRUE,
log = NULL,
...
)Arguments
- object1
First
MizerParamsorMizerSimobject.- object2
Second
MizerParamsorMizerSimobject.- name1, name2
Labels for the two objects, used in the linetype legend.
- power
The abundance is plotted as the number density times the weight raised to
power. The defaultpower = 1gives the biomass density, whereaspower = 2gives the biomass density with respect to logarithmic size bins.- log_x
If
TRUE(default), use a log10 x-axis.- log_y
If
TRUE(default), use a log10 y-axis.- log
Character string specifying which axes should use log10 scales, in the same form as the base
plot()argument. For example,"x","y","xy"or"". If supplied, this overrideslog_xandlog_y.- ...
Arguments passed to
plotSpectra()for preparing the spectra data, for examplespecies,time_range,wlim,ylim,resource,backgroundortotal.
See also
Other plotting functions:
addPlot(),
animate.ArrayTimeBySpeciesBySize(),
plot,
plot2(),
plotBiomass(),
plotCDF(),
plotCDF2(),
plotDiet(),
plotFMort(),
plotFeedingLevel(),
plotGrowthCurves(),
plotMizerParams,
plotMizerSim,
plotPredMort(),
plotRelative(),
plotSpectra(),
plotSpectraRelative(),
plotYield(),
plotYieldGear(),
plotting_functions