plotCDF() plots the cumulative distribution over body size from small to
large sizes. It uses the same spectra data preparation as plotSpectra().
The density is first multiplied by w^power, then integrated over size.
With normalise = TRUE, each curve is divided by its final value so that it
ends at 1.
Usage
plotCDF(object, ...)
# S3 method for class 'MizerSim'
plotCDF(
object,
species = NULL,
time_range,
geometric_mean = FALSE,
wlim = c(NA, NA),
ylim = c(NA, NA),
power = 1,
biomass = TRUE,
total = FALSE,
resource = FALSE,
background = TRUE,
highlight = NULL,
normalise = TRUE,
log_x = TRUE,
log = NULL,
return_data = FALSE,
...
)
# S3 method for class 'MizerParams'
plotCDF(
object,
species = NULL,
wlim = c(NA, NA),
ylim = c(NA, NA),
power = 1,
biomass = TRUE,
total = FALSE,
resource = FALSE,
background = TRUE,
highlight = NULL,
normalise = TRUE,
log_x = TRUE,
log = NULL,
return_data = FALSE,
...
)
plotlyCDF(
object,
species = NULL,
time_range,
geometric_mean = FALSE,
wlim = c(NA, NA),
ylim = c(NA, NA),
power = 1,
biomass = TRUE,
total = FALSE,
resource = FALSE,
background = TRUE,
highlight = NULL,
normalise = TRUE,
log_x = TRUE,
log = NULL,
...
)Arguments
- object
An object of class MizerSim or MizerParams.
- ...
Other arguments (currently unused)
- species
The species to be selected. Optional. By default all target species are selected. A vector of species names, or a numeric vector with the species indices, or a logical vector indicating for each species whether it is to be selected (TRUE) or not.
- time_range
The time range (either a vector of values, a vector of min and max time, or a single value) to average the abundances over. Default is the final time step. Ignored when called with a MizerParams object.
- geometric_mean
If TRUE then the average of the abundances over the time range is a geometric mean instead of the default arithmetic mean.
- wlim
A numeric vector of length two providing lower and upper limits for the w axis. Use NA for the default: the lower default is
min(params@w) / 100whenresource = TRUE(to show some resource below the fish grid) ormin(params@w)whenresource = FALSE; the upper default ismax(params@w_full). Data is filtered to this range and the axis limits are set accordingly.- ylim
A numeric vector of length two providing lower and upper limits for the y axis. Use NA to auto-scale to the data range. Values below 1e-20 are always filtered out from the data regardless of
ylim[1]. Data aboveylim[2]is filtered and the upper axis limit is set accordingly.- power
The abundance is plotted as the number density times the weight raised to
power. The defaultpower = 1gives the biomass density, whereaspower = 2gives the biomass density with respect to logarithmic size bins.- biomass
Only used if
powerargument is missing. Thenbiomass = TRUEis equivalent topower=1andbiomass = FALSEis equivalent topower=0- total
A boolean value that determines whether the total over all species in the system is plotted as well. Note that even if the plot only shows a selection of species, the total is including all species. Default is FALSE.
- resource
A boolean value that determines whether resource is included. Default is FALSE.
- background
A boolean value that determines whether background species are included. Ignored if the model does not contain background species. Default is TRUE.
- highlight
Name or vector of names of the species to be highlighted.
- normalise
If
TRUE(default), plot the cumulative proportion. IfFALSE, plot the cumulative abundance, biomass, or other unnormalised integral.- log_x
If
TRUE(default), use a log10 x-axis.- log
Character string specifying whether the x-axis should use a log10 scale, in the same form as the base
plot()argument. ForplotCDF(), only"x"and""are supported. If supplied, this overrideslog_x.- return_data
A boolean value that determines whether the formatted data used for the plot is returned instead of the plot itself. Default value is FALSE
Value
A ggplot2 object, unless return_data = TRUE, in which case a data
frame with the four variables 'w', 'value', 'Species', 'Legend' is
returned.
plotlyCDF() returns a plotly object.
See also
Other plotting functions:
addPlot(),
animate.ArrayTimeBySpeciesBySize(),
plot,
plot2(),
plotBiomass(),
plotCDF2(),
plotDiet(),
plotFMort(),
plotFeedingLevel(),
plotGrowthCurves(),
plotMizerParams,
plotMizerSim,
plotPredMort(),
plotRelative(),
plotSpectra(),
plotSpectra2(),
plotSpectraRelative(),
plotYield(),
plotYieldGear(),
plotting_functions
Examples
# \donttest{
plotCDF(NS_params, species = c("Cod", "Herring"))
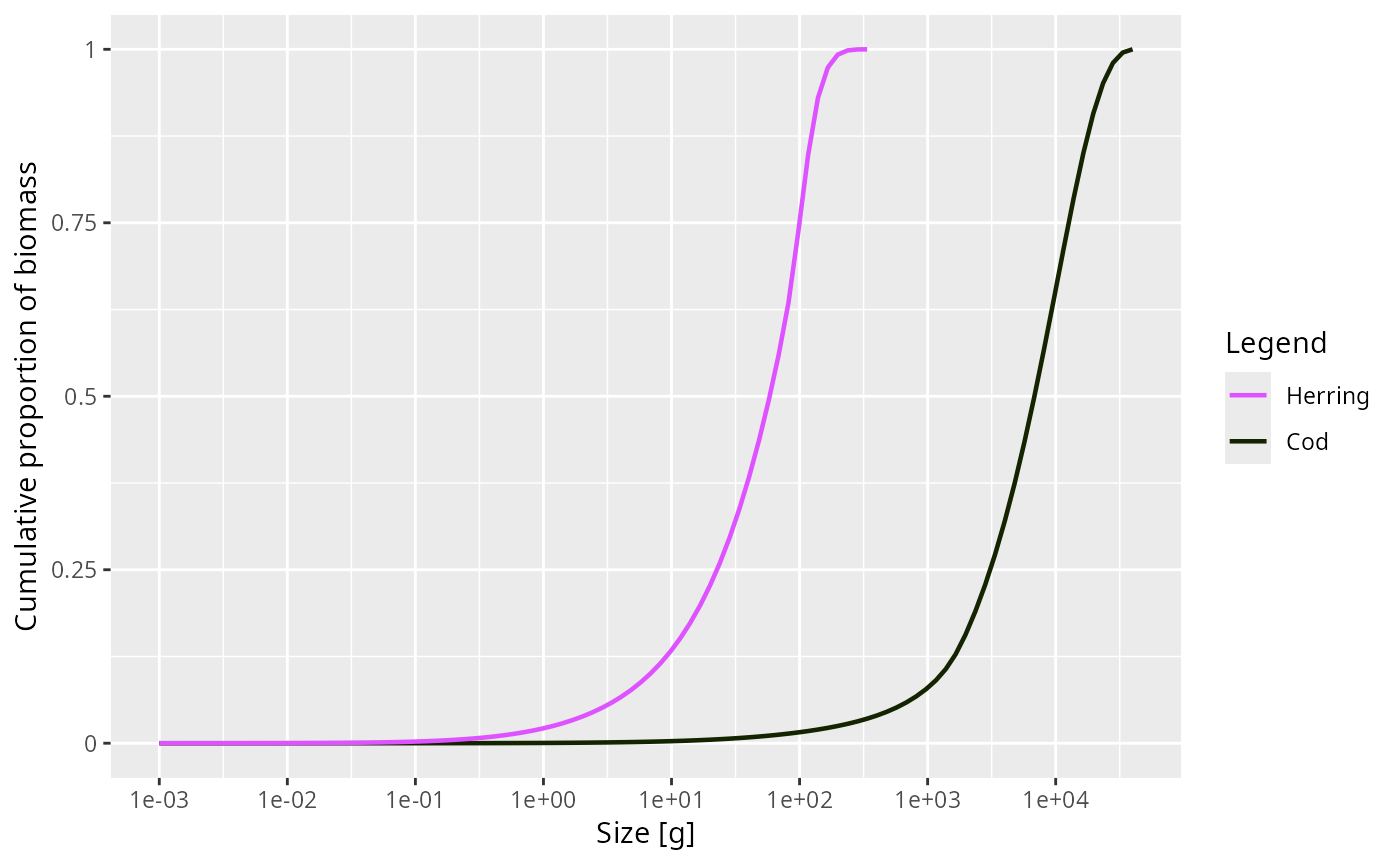 plotCDF(NS_sim, power = 0, normalise = FALSE)
plotCDF(NS_sim, power = 0, normalise = FALSE)
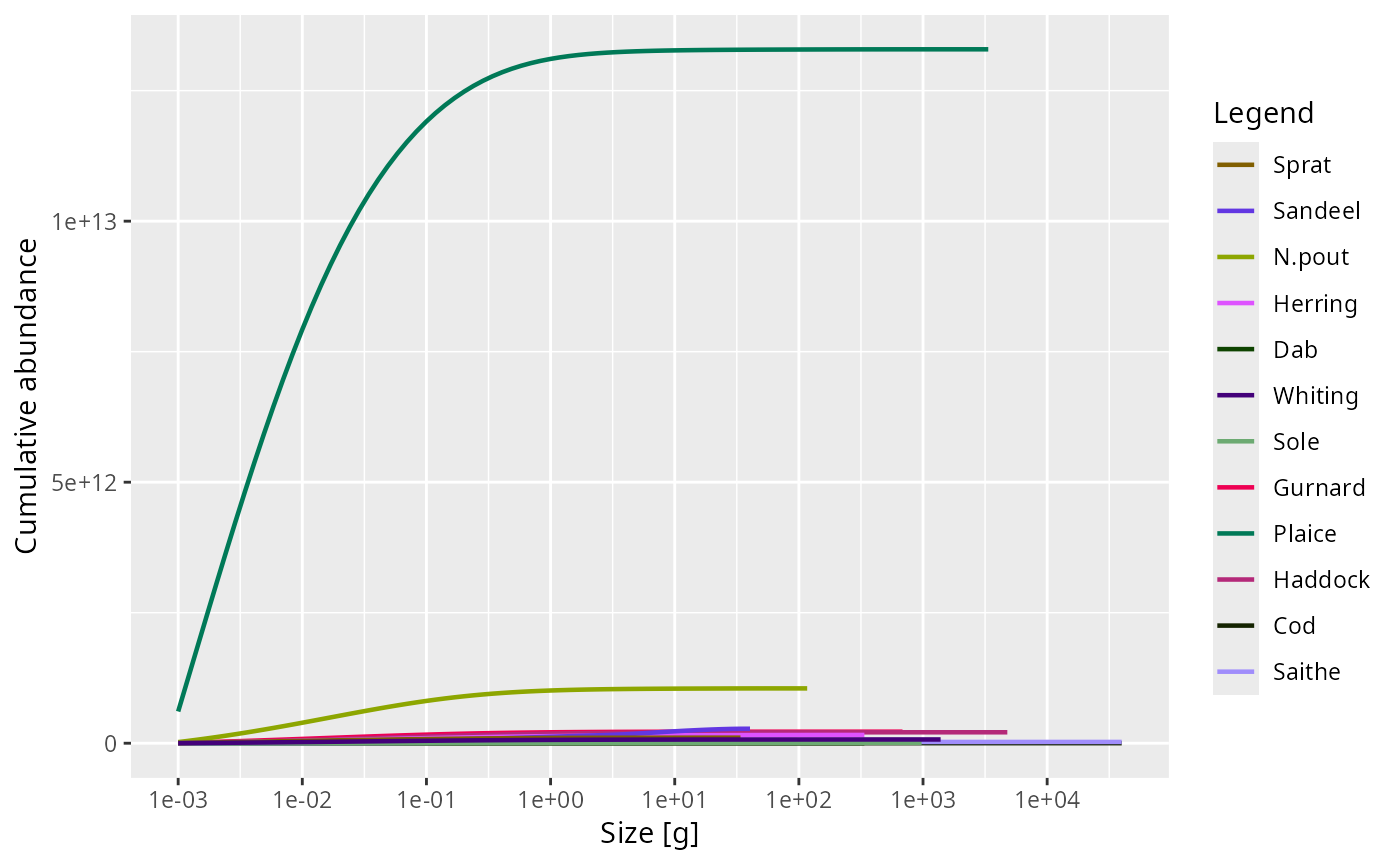 # }
# }
