
Compare two mizer array objects in one plot
Source:R/ArraySpeciesBySize-class.R, R/ArrayTimeBySpecies-class.R, R/ArrayTimeBySpeciesBySize-class.R
plot2.Rdplot2() compares two compatible mizer array objects in a single ggplot.
Colours identify species or groups, and linetype identifies which object the
values came from.
Usage
plot2(x, y, ...)
# S3 method for class 'ArraySpeciesBySize'
plot2(
x,
y,
name1 = "First",
name2 = "Second",
species = NULL,
all.sizes = FALSE,
log_x = TRUE,
log_y = FALSE,
log = NULL,
wlim = c(NA, NA),
ylim = c(NA, NA),
total = FALSE,
background = TRUE,
y_ticks = 6,
...
)
# S3 method for class 'ArrayTimeBySpecies'
plot2(
x,
y,
name1 = "First",
name2 = "Second",
species = NULL,
start_time = NULL,
end_time = NULL,
y_ticks = 6,
ylim = c(NA, NA),
total = FALSE,
background = TRUE,
log_x = FALSE,
log_y = TRUE,
log = NULL,
...
)
# S3 method for class 'ArrayTimeBySpeciesBySize'
plot2(
x,
y,
name1 = "First",
name2 = "Second",
species = NULL,
time = NULL,
all.sizes = FALSE,
log_x = TRUE,
log_y = FALSE,
log = NULL,
wlim = c(NA, NA),
ylim = c(NA, NA),
total = FALSE,
background = TRUE,
y_ticks = 6,
...
)Arguments
- x, y
Two compatible mizer array objects of the same class.
- ...
Further arguments (currently unused).
- name1, name2
Labels for the two objects, used in the linetype legend.
- species
Character vector of species to include.
NULL(default) means all species.- all.sizes
If
FALSE(default), values outside a species' size range (w_mintow_max) are removed. Only applies toArraySpeciesBySize.- log_x
If
TRUE, use a log10 x-axis. Default isTRUEfor size spectra andFALSEfor time series.- log_y
If
TRUE, use a log10 y-axis. Default isFALSEforArraySpeciesBySizeandTRUEforArrayTimeBySpecies.- log
Character string specifying which axes should use log10 scales, in the same form as the base
plot()argument. For example,"x","y","xy"or"". If supplied, this overrideslog_xandlog_y.- wlim
A numeric vector of length two providing lower and upper limits for the weight (x) axis. Use
NAto refer to the existing minimum or maximum. Only applies toArraySpeciesBySize.- ylim
A numeric vector of length two providing lower and upper limits for the value (y) axis. Use
NAto refer to the existing minimum or maximum.- total
A boolean value that determines whether the total over all selected species is plotted as well. Default is
FALSE.- background
A boolean value that determines whether background species are included. Ignored if the model does not contain background species. Default is
TRUE.- y_ticks
The approximate number of ticks desired on the y axis.
- start_time
The first time to be plotted. Default (
NULL) is the beginning of the time series. Only applies toArrayTimeBySpecies.- end_time
The last time to be plotted. Default (
NULL) is the end of the time series. Only applies toArrayTimeBySpecies.- time
The time to display. Default (
NULL) is the final time step. Only applies toArrayTimeBySpeciesBySize.
See also
Other plotting functions:
addPlot(),
animate.ArrayTimeBySpeciesBySize(),
plot,
plotBiomass(),
plotCDF(),
plotCDF2(),
plotDiet(),
plotFMort(),
plotFeedingLevel(),
plotGrowthCurves(),
plotMizerParams,
plotMizerSim,
plotPredMort(),
plotRelative(),
plotSpectra(),
plotSpectra2(),
plotSpectraRelative(),
plotYield(),
plotYieldGear(),
plotting_functions
Examples
# \donttest{
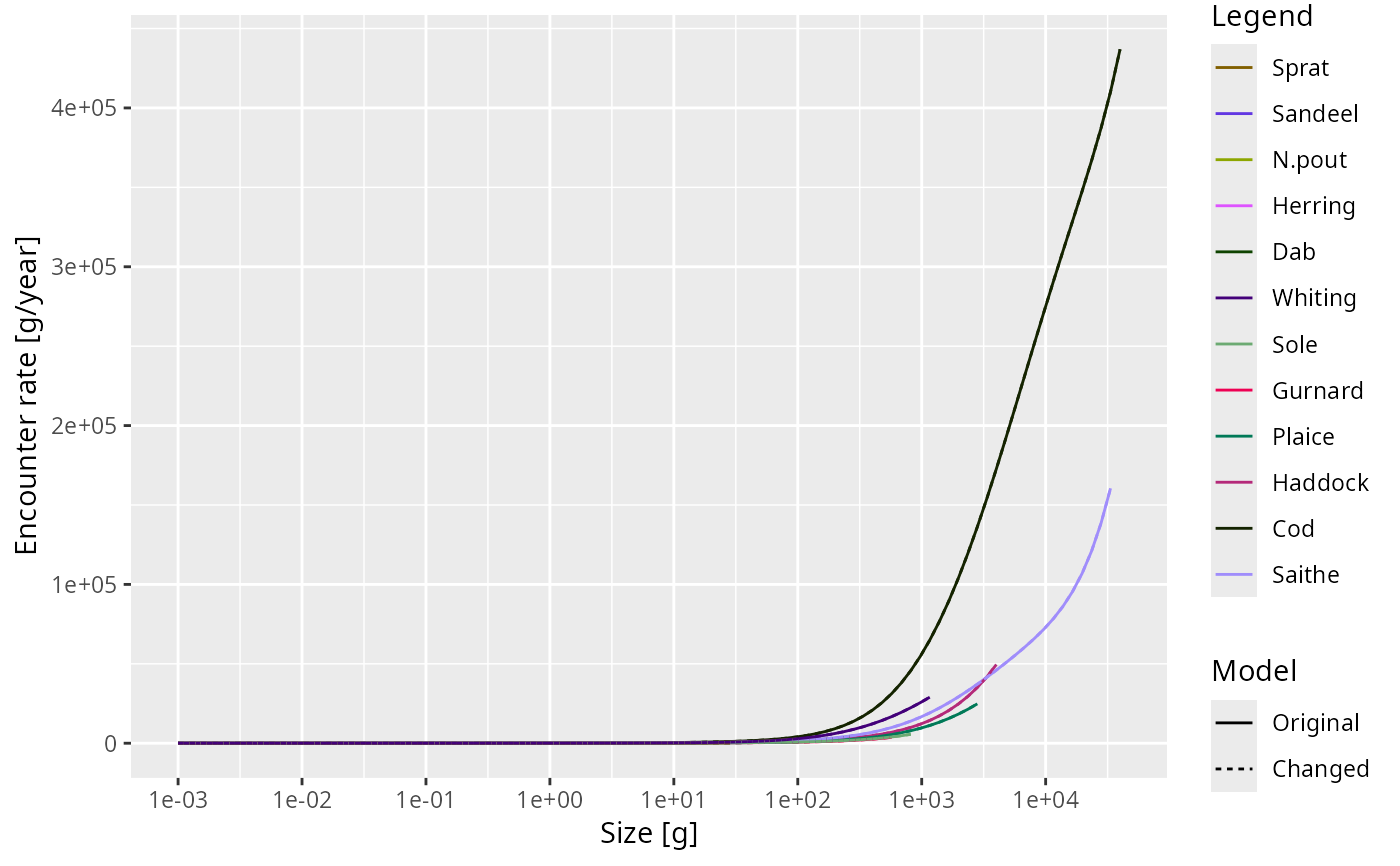
enc <- getEncounter(NS_params)
plot2(enc, enc, name1 = "Original", name2 = "Changed")
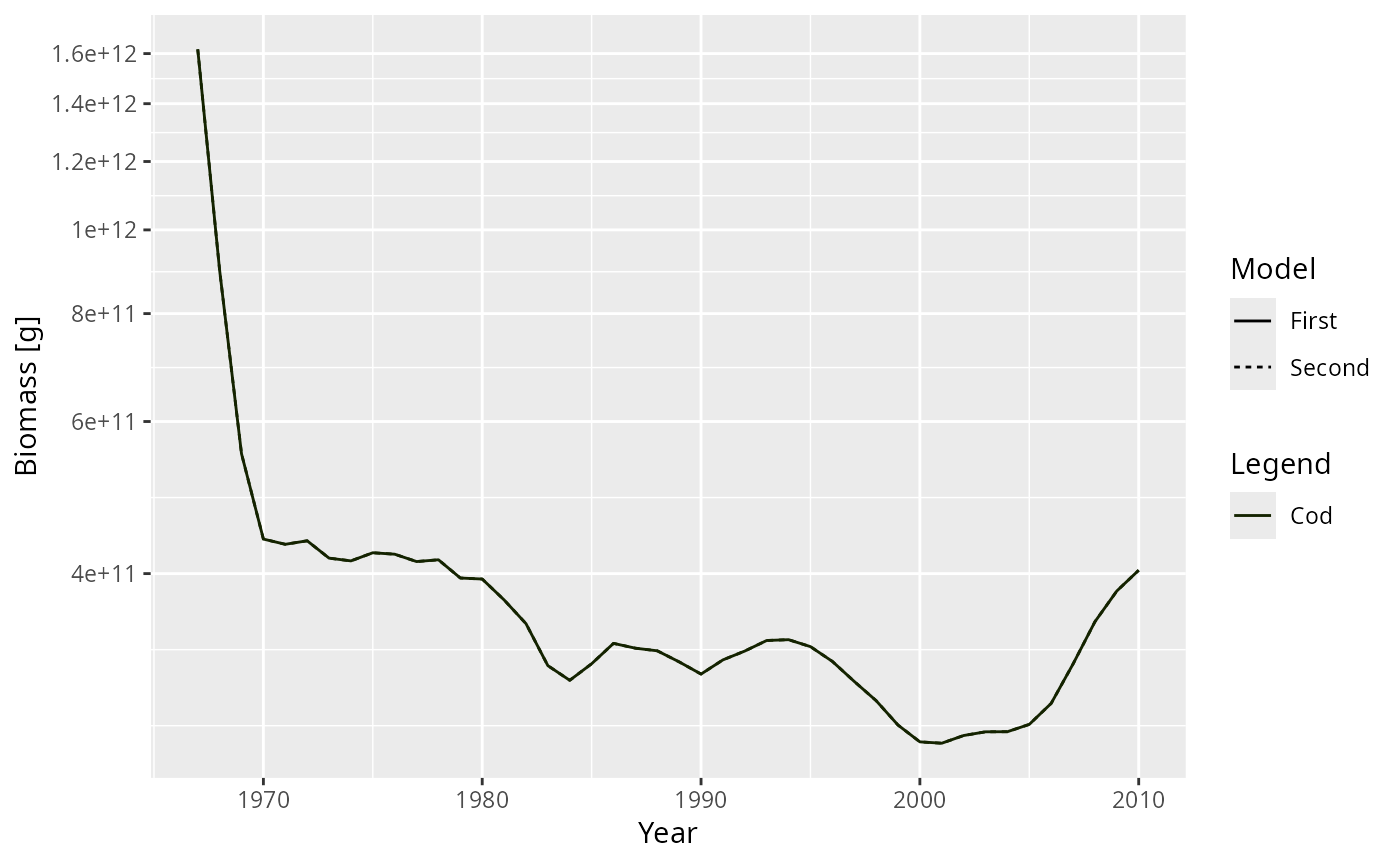
 plot2(getBiomass(NS_sim), getBiomass(NS_sim), species = "Cod")
plot2(getBiomass(NS_sim), getBiomass(NS_sim), species = "Cod")
 # }
# }