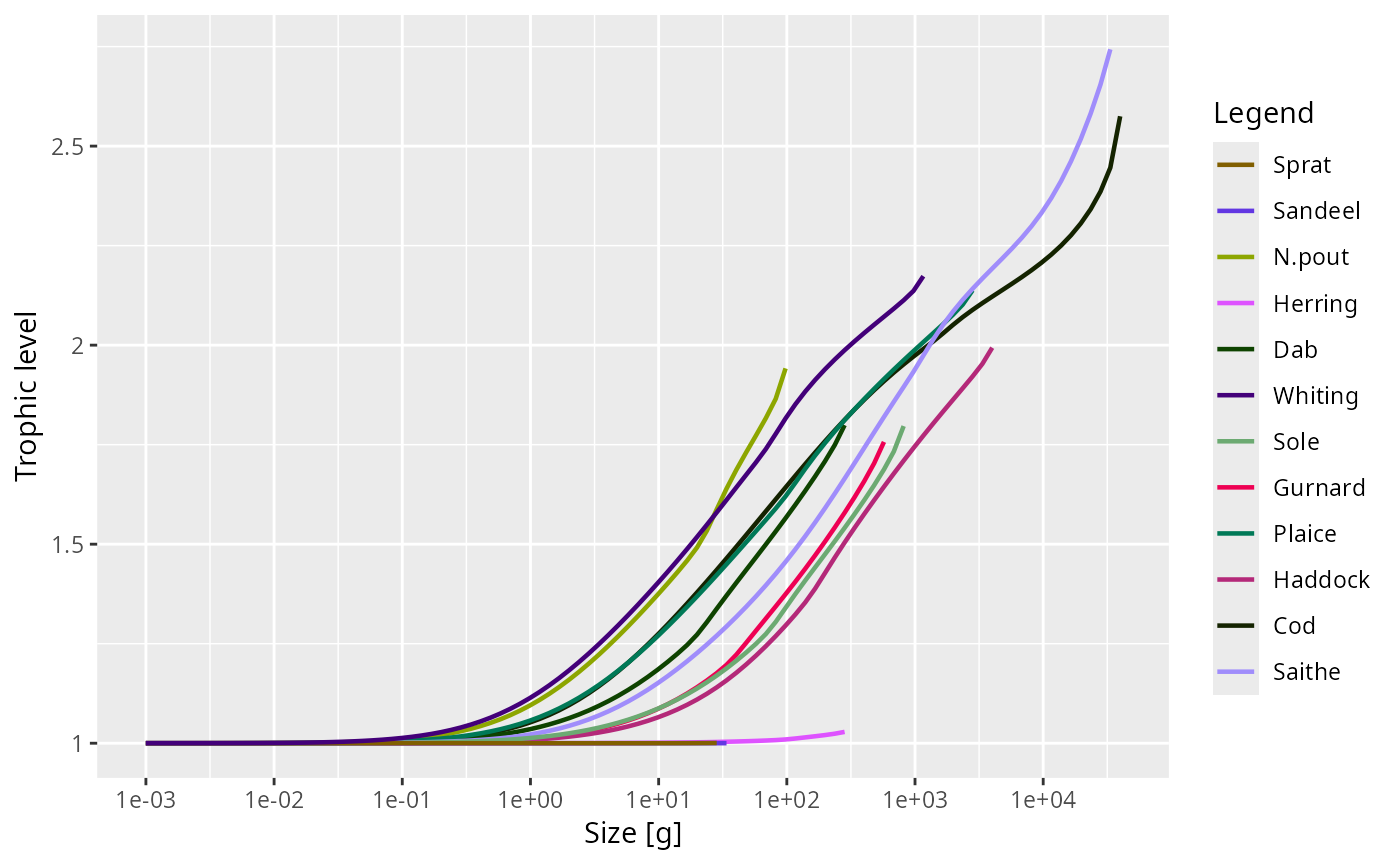
Calculates the trophic level of individuals of each species at each size,
assuming the system is in a steady state. The trophic level of an individual
is defined as 1 more than the consumption-rate-weighted average trophic level
of all the prey it has consumed during its lifetime up to the current size.
The trophic level of the primary resource is set to 0.
Usage
getTrophicLevel(
params,
n = initialN(params),
n_pp = initialNResource(params),
n_other = initialNOther(params),
...
)Arguments
- params
A MizerParams object
- n
A matrix of species abundances (species x size).
- n_pp
A vector of the resource abundance by size
- n_other
A list of abundances for other dynamical components of the ecosystem
- ...
Unused
Value
An ArraySpeciesBySize object (species x size) with the trophic
level of individuals at each size. Entries below the egg size of each
species are NA.
Details
In the traditional non-size-resolved approach, all individuals of a species have the same diet composition \(D_{ij}\), defined as the proportion of total biomass intake of species \(i\) that comes from species \(j\). The trophic levels then satisfy $$T_i = 1 + \sum_j D_{ij}\,T_j,$$ which is solved as a linear system \((I - D)\,\mathbf{T} = \mathbf{1}\).
In mizer, diet composition changes as an individual grows, so we must integrate over the individual's lifetime. Assuming a steady state so that the growth rate \(g_i(w)\) and prey densities depend only on size and not on time, we can replace the integral over time since birth by an integral over weight using \(dt = dw / g_i(w)\). The trophic level \(T_i(w)\) of an individual of species \(i\) at weight \(w\) is then $$ T_i(w) = 1 + \frac{ \int_{w_0}^{w} \frac{1}{g_i(w')} \sum_j \int r_{ij}(w', w_p)\, T_j(w_p)\, dw_p\, dw' }{ \int_{w_0}^{w} \frac{1}{g_i(w')} \sum_j \int r_{ij}(w', w_p)\, dw_p\, dw' }, $$ where \(w_0\) is the egg size and \(r_{ij}(w, w_p)\) is the rate at which a predator of species \(i\) at weight \(w\) consumes biomass from prey species \(j\) at weight \(w_p\): $$ r_{ij}(w, w_p) = \theta_{ij}\,\gamma_i(w)\,(1 - f_i(w))\,\phi_i(w/w_p)\, N_j(w_p)\,w_p. $$ The sum over \(j\) runs over all species. The resource is excluded from the numerator because its trophic level is 0, but is included in the denominator (which equals the total biomass consumed over the predator's lifetime from egg size to current weight \(w\)).
This equation can be viewed as a linear system \((I - D)\,\mathbf{T} = \mathbf{1}\) in which the entries of \(\mathbf{T}\) are indexed by \((i, w)\) and the matrix \(D\) encodes the lifetime-integrated diet composition. The system is solved iteratively from small to large sizes, exploiting the fact that prey are typically much smaller than the predator (large predator-to-prey mass ratio), so that the trophic levels of all relevant prey sizes are already known when computing \(T_i(w)\).
See also
Other summary functions:
getBiomass(),
getDiet(),
getGrowthCurves(),
getN(),
getSSB(),
getTrophicLevelBySpecies(),
getYield(),
getYieldGear()
Examples
tl <- getTrophicLevel(NS_params)
plot(tl)