This function has been deprecated and will be removed in the future unless
you have a use case for it. If you do have a use case for it, please let the
developers know by creating an issue at
https://github.com/sizespectrum/mizer/issues.
Arguments
- params
A MizerParams object
- species
The species to be affected. Optional. By default all observed yields will be matched. A vector of species names, or a numeric vector with the species indices, or a logical vector indicating for each species whether it is to be affected (TRUE) or not.
- info_level
Controls the amount of information messages that are shown. Higher levels lead to more messages.
- ...
Additional arguments passed to the method.
Details
If you want to match the yields to observations, you should use the matchYield() function from the mizerExperimental package instead, which adjusts the catchability to match the yield rather than by adjusting the biomass.
The function adjusts the abundances of the species in the model so that their yearly yields under the given fishing mortalities match with observations.
The function works by multiplying for each species the abundance density
at all sizes by the same factor. This will of course not give a steady
state solution, even if the initial abundance densities were at steady state.
So after using this function you may want to use steady() to run the model
to steady state, after which of course the yields will no longer match
exactly. You could then iterate this process. This is described in the
blog post at https://blog.mizer.sizespectrum.org/posts/2021-08-20-a-5-step-recipe-for-tuning-the-model-steady-state/.
Before you can use this function you will need to have added a
yield_observed column to your model which gives the observed yields in
grams per year. For species for which you have no observed biomass, you
should set the value in the yield_observed column to 0 or NA.
Examples
params <- NS_params
species_params(params)$yield_observed <-
c(0.8, 61, 12, 35, 1.6, 20, 10, 7.6, 135, 60, 30, 78)
gear_params(params)$catchability <-
c(1.3, 0.065, 0.31, 0.18, 0.98, 0.24, 0.37, 0.46, 0.18, 0.30, 0.27, 0.39)
params <- calibrateYield(params)
params <- matchYields(params)
#> The following species are not being fished in your model and their abundances will not be changed: Sprat, Sandeel, N.pout.
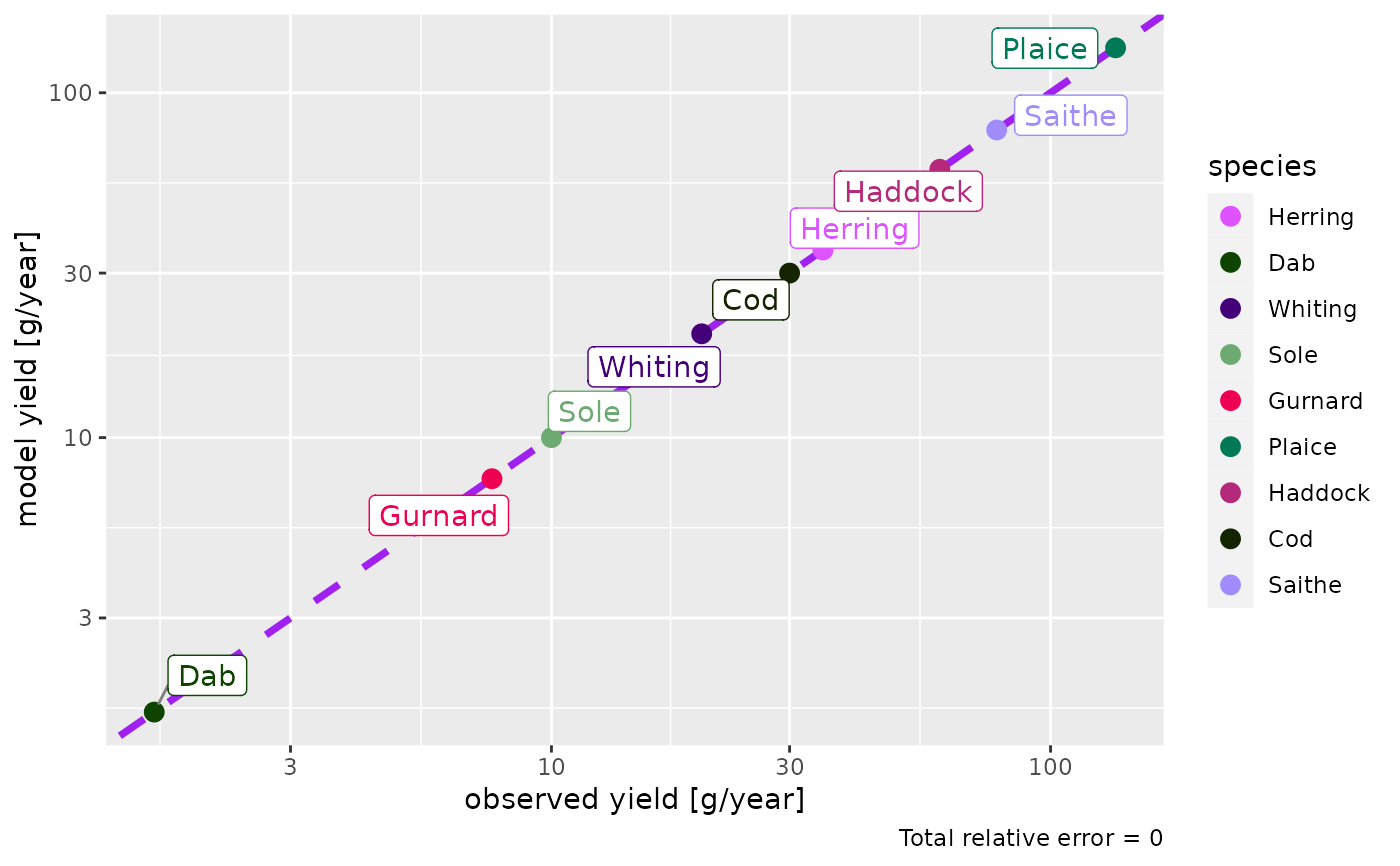
plotYieldObservedVsModel(params)
#> The following species are not being fished in your model and will not be included in the plot: Sprat, Sandeel, N.pout.